Science
RESEARCH FOCUS
One of the key aims of the BCRB is to better understand the mechanisms that tightly control the redox homeostasis of the cell under stress conditions. We are studying the mechanisms behind reactive oxygen species scavenging, signaling and regulation. Several oxidoreductase proteins, which successively pass on electrons via complex intra- and intermolecular cascades using thiol-disulfide chemistry, are involved. We specialize in the in vitro reconstitution of these thiol/disulfide electron transfer pathways, the study of the kinetics and structural changes during electron transfer, and the discovery of new cellular pathways involved in redox signaling.
• A long-term objective of the Collet lab is to better understand the redox mechanisms that govern the assembly of the cell envelope and maintain its integrity during the cell cycle.
• The mission of the Messens lab is to decipher how cells sense redox metabolites and transduce stimuli into downstream biological effects.
• The mission of the Van Breusegem lab is to study the impact and the molecular response of plants against increased Reactive Oxygen Species levels.
The coming years we want to exploit our expertise in thiol/redox-biochemistry through the following projects:
• ‘Redox signaling networks in plants'
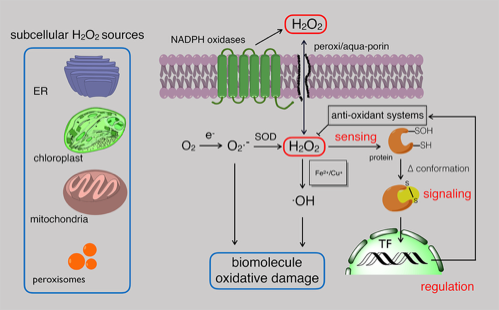
ROS has emerged as important regulator of plant stress responses. Perturbation in ROS production and/or scavenging are sensed by plant cells as a 'warning' message, and genetic programs leading to stress acclimation or cell death are switched on. Knowledge on regulatory events during hydrogen peroxide signal transduction is now only scratching the surface. Through combined top-down and bottom-up genomics and proteomics approaches the Van Breusegem and Messens labs are dissecting the network governing H2O2 signal transduction in plants and pinpoint genes that are potential candidates for innovative molecular breeding strategies to develop stress-tolerant crops.
• ‘Understanding the redox pathways involved in protein folding’
How proteins fold in the cell envelope is a long-standing mystery. Several protein folding catalysts and chaperones have been identified in the envelope, but how they assist protein folding is only partially understood. We and others have significantly contributed to the characterisation of the pathways that oxidatively fold envelope proteins in E. coli. The Collet lab now wants to address the intriguing problems that are still unsolved regarding the oxidative protein folding.
• 'Thiol-based redox signaling'
Today, we know that H2O2 can initiate various signaling cascades, but a long way remains towards a fully-characterized H2O2 signaling network. Full characterization requires more information regarding H2O2 perception, transduction, and regulation as well as knowledge of how this network intersects with other signaling networks such as those involving Ca2+ or phosphorylation. The Messens lab aims to understand the structural and chemical microenvironment within redox-sensing proteins that determines the kinetic privilege of cysteines to react with H2O2.
• 'Peroxide signaling facilitators'
To address the question of H2O2 signal transduction and regulation, we will be investigating peroxiredoxins (Prdx). Prdx are highly efficient and highly abundant peroxide-scavenging enzymes, which have recently also been demonstrated to participate in redox-based protein-protein interactions with the possibility to transferring the oxidizing peroxide signal to other proteins that are less reactive towards oxidation byH2O2. One of the objectives of the Messens lab is to explore the hypothesized protein scaffolding system which engenders specificity in this Prdx-mediated peroxide signaling.
• 'Metabolite indicator tools'
In order to be able to investigate cellular (redox) metabolite signaling between organelles and cells it is important to have proper tools enabling the visualization of metabolic fluxes in real time in living cells, with a subcellular resolution which mass spectrometry “omics” methods cannot afford. In response to this need, the Messens lab developing metabolite indicators which harness metabolite-responsive proteins to create sensitive and specific biosensors with a fluorescence read-out that could be used in various cell types and organisms.
• ‘Discovering the molecular mechanisms activating the Rcs cascade’
The Collet lab wants to understand how the Rcs phosphorelay is activated. The Rcs pathway, which is one of the major envelope stress response systems in E. coli, controls the expression of genes involved in the protection of envelope integrity, in biofilm formation and in virulence-associated structures.
3